GoodWorkflows¶
A DSL2 Nextflow pipeline for composing reusable single-cell RNA-seq workflows and running them on SLURM + Podman HPC systems.
Workflow selection guide¶
| Workflow | What it does | Compute requirement |
|---|---|---|
full |
Download → export counts → harmonize → scMODAL integration | HPC + GPU (SLURM required) |
ingest_export |
Download Seurat RDS and export 10x-like counts | Local / Mac / HPC (CPU) |
ingest_tabulate |
Download cell metadata and build subjectIdTable.csv |
Local / Mac / HPC (CPU) |
Select the workflow with --workflow <name>.
Quick start¶
Local / Mac (CPU workflows)¶
# Download metadata and produce a subject-level summary table
nextflow run main.nf \
--workflow ingest_tabulate \
--labkey_base_url https://labkey.example.org \
--labkey_folder /My/Project/Folder
# Download Seurat objects and export 10x-like counts
nextflow run main.nf \
--workflow ingest_export \
--labkey_base_url https://labkey.example.org \
--labkey_folder /My/Project/Folder
HPC (full GPU pipeline)¶
# On any SLURM node – profile auto-detected from environment
sbatch slurm_nextflow.sh \
--workflow full \
--labkey_base_url https://labkey.example.org \
--labkey_folder /My/Project/Folder
LabKey credentials
All workflows communicate with a LabKey / Prime-seq server via ~/.netrc. Ensure your netrc
entry for labkey_base_url is configured before running.
Repository layout¶
.
├── main.nf # Thin launcher
├── workflows/ # Higher-level workflow definitions
├── modules/local/ # Single-step DSL2 modules
├── configs/ # Base + profile-specific configs
├── data/ # Default input location (samplesheet.csv)
├── outputs/ # Published results (generated)
├── work/ # Nextflow work dir (generated)
├── logs/ # Nextflow reports and SLURM logs (generated)
├── docs/ # Documentation source (this site)
├── mkdocs.yml # MkDocs site config
├── nextflow_schema.json # Machine-readable parameter API (JSON Schema)
├── slurm_nextflow.sh # HPC SLURM submission wrapper
└── slurm_sync_repo.sh # Lightweight HPC repo sync job
Representative outputs¶
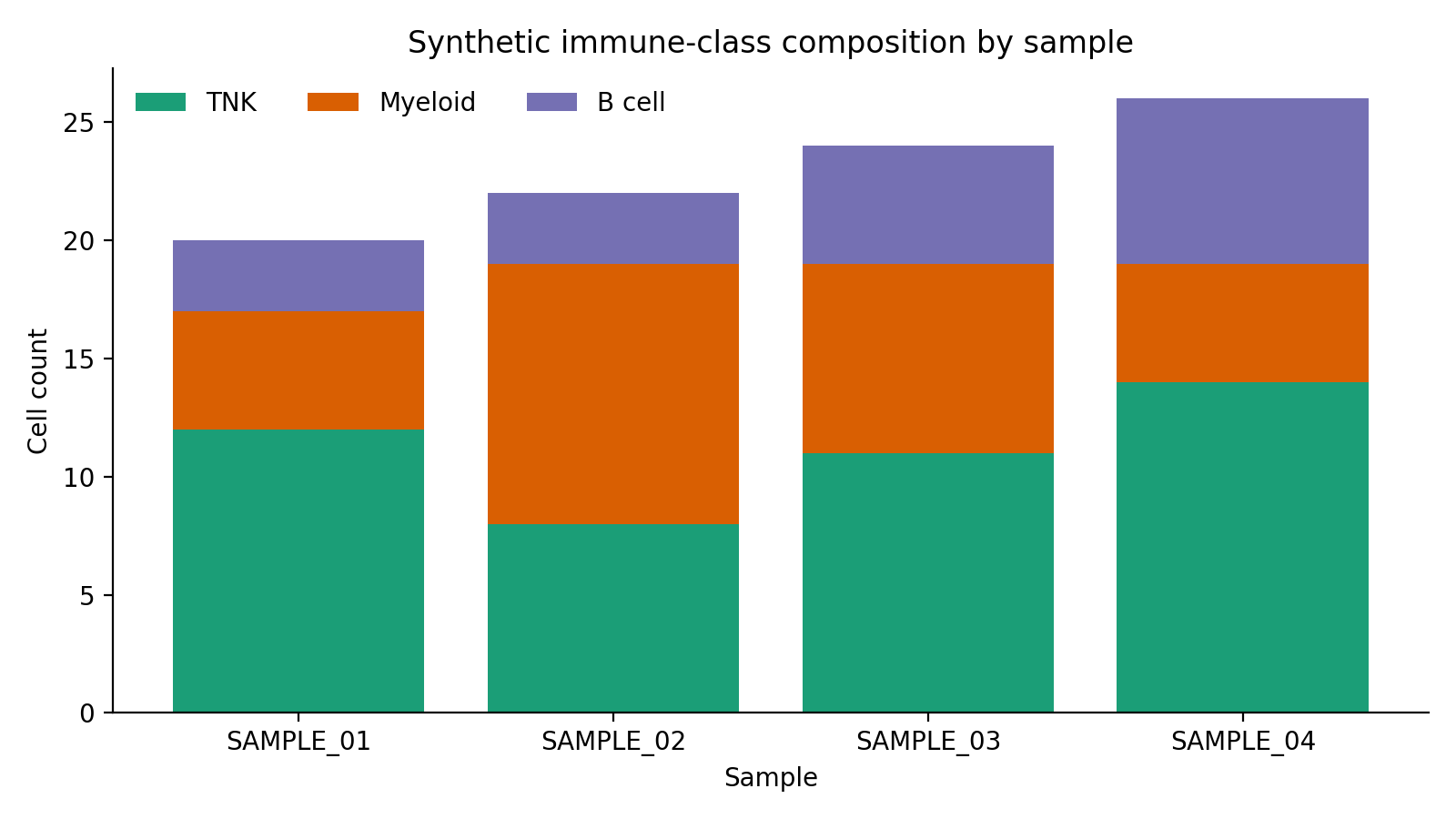
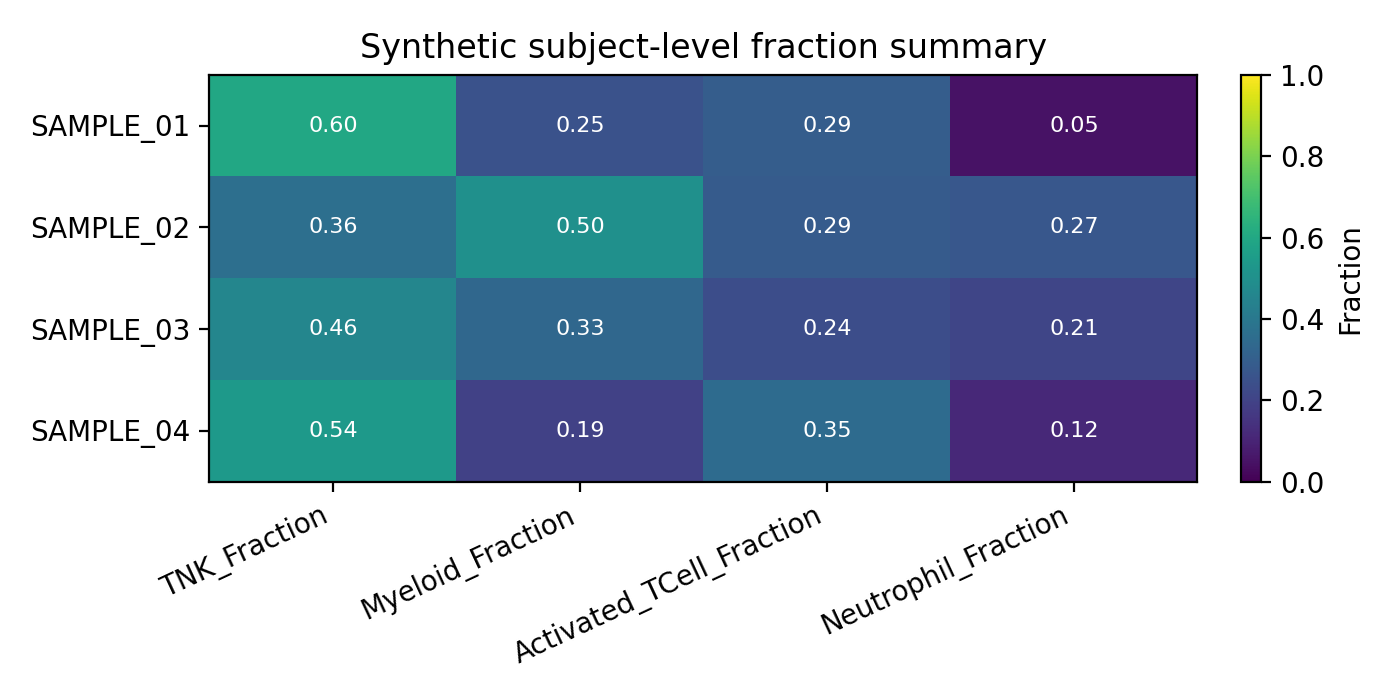
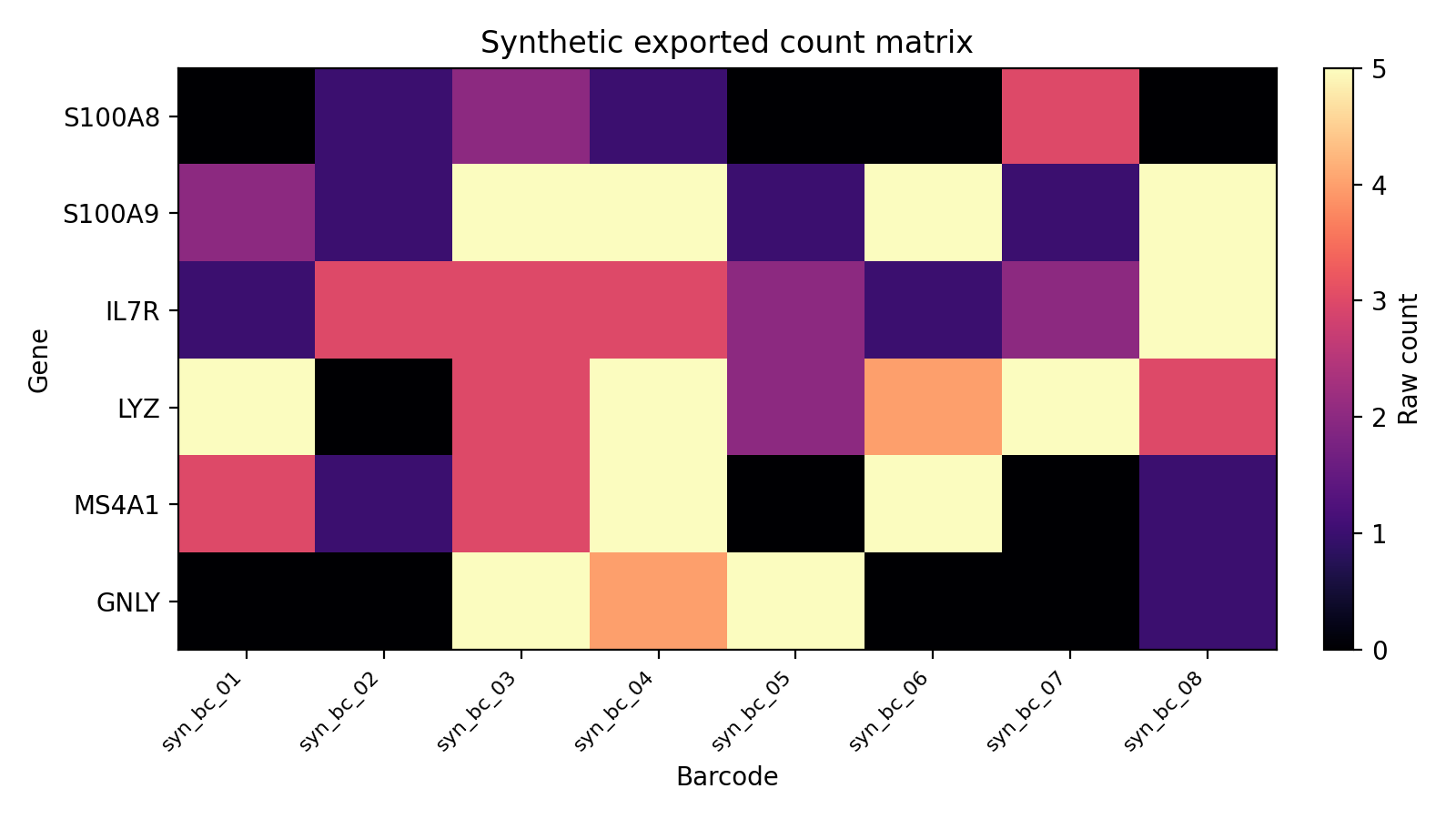
The docs site ships with seeded synthetic examples so the workflow pages can show safe, reproducible output shapes without live LabKey access.
| Cell metadata composition | Subject-level tabulation | Exported count matrix |
|---|---|---|
 |
 |
 |
See the Synthetic Tabulation Walkthrough for the full end-to-end explanation of where these files come from and how they map to the workflows.
Generated API reference¶
The API Reference section is rebuilt with uvx nf-docs generate during docs CI. Use those pages for code- and schema-level reference, and use the curated workflow pages for stage-by-stage semantics, expected file layouts, and visual examples.